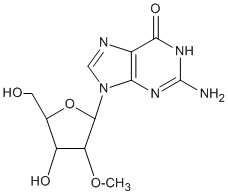
Synthetic oligonucleotides, just like their natural counterparts, are prone to degradation once introduced into a cell. This degradation is due to the presence of exo and endonuclease enzymes, as well as inherent chemical instability (particularly for RNA). Under cellular conditions, this leads to fast in vivo degradation of oligos and a short half-life. (1) To reduce or eliminate this susceptibility, nuclease-resistant modifications can be introduced into oligonucleotides. For antisense or RNAi applications, incorporation of modifications conferring nuclease resistance is essential and such modifications are used routinely. There are a number of ways to introduce nuclease resistance into a synthetic oligonucleotide. Most commonly, the substitution of 2'-OMe bases at some or all positions of an oligo is used as the preferred route to inducing nuclease resistance.(2) Since the nuclease resistance conferred by 2'-OMe lies between that of unmodified nucleosides (no resistance) and phosphorothiolation (highly resistant), extensive/complete 2'-O-methylation is frequently chosen when a high level of nuclease resistance is required. 2'-O-methylation also confers the desirable property of higher binding affinity (that is, higher duplex Tm) to the oligo for its target. For these reasons, 2'-OMe nucleosides are extensively used in siRNA and aptamer applications. 2'-O-Methyloligoribonucleotides are extremely useful reagents for a variety of molecular biology applications. The 2'-OMe RNA-RNA duplex is more thermally stable than the corresponding DNA-RNA one.(3) In addition, 2'-OMe-RNA is chemically more stable than either DNA or RNA and is resistant to degradation by RNA- or DNA-specific nucleases.(4) It is worth noting though that duplexes formed between oligos having 2'-OMe bases at all positions and RNA are incapable of RNase H activity, thus making them ineffective in RNaseH dependent antisense applications,(5) although they can suppress gene expression by blocking the mRNA translation process via steric hindrance.(6) We provide a range of 2'-OMe CPGs with a variety of pore sizes and linkers consistent with our unmodified DNA and RNA CPG products. The protecting group strategies are compatible with the usual DNA and RNA chemistries. Note that the 2'-OMe group in itself is not a 2'-OH protecting group strategy; the 2'-OMe group cannot be cleaved under RNA synthesis and deprotection conditions. Depending on the product, various column types (Standard and ALL-FIT luer, and MerMade, Super and Hybrid pipette columns).
Ref:
- Rate of degradation of {alpha} and {beta}-oligodeoxynucleotides in Xenopus oocytes. Implications for anti-messenger strategies, C. Cazenave, M. Chevrier, T.T. Nguyen and C. Helene, Nucleic Acids Research, 15, 10507- 10521, 1987.
- (a) Evaluation of 2'-Modified Oligonucleotides Containing 2'-Deoxy Gaps as Antisense Inhibitors of Gene Expression, B.P. Monia, E.A. Lesnik, C. Gonzalez, W.F. Lima, D. McGee, C.J. Guinosso, A.M. Kawasaki, P.D. Cook and S.M. Frier, J. Biol. Chem., 268, 14514-14522, 1993; (b) Nuclease Resistance and Antisense Activity of Modified Oligonucleotides Targeted to Ha-ras, B.P. Monia, J.F. Johnston, H. Sasmor and L.L. Cummins, J. Biol. Chem., 271, 14533-14540, 1996.
- Synthesis and hybridization studies on two complementary nona(2'-O-methyl)ribonucleotides, H. Inoue, Y. Hayase, A. Imura, S. Iwai, K. Miura, and E. Ohtsuka, Nucleic Acids Research, 15, 6131-6148, 1987.
- Highly efficient chemical synthesis of 2'-O-methyloligoribonucleotides and tetrabiotinylated derivatives; novel probes that are resistant to degradation by RNA or DNA specific nucleases, B.S. Sproat, A.I. Lamond, B. Beijer, P. Neuner and U. Ryder, Nucleic Acids Research, 17, 3373-3386, 1989.
- Sequence-dependent hydrolysis of RNA using modified oligonucleotide splints and RNase H, H. Inoue, Y. Hayase, S. Iwai and E. Ohtsuka, FEBS Lett., 215, 327-330, 1987.
- Antisense technologies. Improvement through novel chemical modifications, J. Kurreck, Eur. J. Biochem., 270, 1628-1644, 2003.
|
Synthesizer
|
Column
|
Type/Description
|
Notes
|
|
MerMade 6,12
|
MerMade, syringe (all scales)
|
Pipette type column
|
A MerMade column is also known as a Supercolumn
|
|
MerMade 4, 48X, 96E, 192E, 192X
|
MerMade, Syringe (up to 1.3 mL)
|
Pipette type column
|
A MerMade column is also known as a Supercolumn
|
|
ABI 384 / 394
|
Luer
|
Barrel column with luer fitting at either end
|
Also known as ALL-FIT or Standard
|
|
Expedite 8909
|
Luer
|
Barrel column with luer fitting at either end
|
Also known as ALL-FIT or Standard
|
|
ABI3900
|
MerMade
|
Pipette type column
|
A MerMade column is also known as a Supercolumn
|
|
K&A H4, H8, H8SE, H2, H32, H64
|
Luer
|
Barrel column with luer fitting at either end
|
Also known as ALL-FIT or Standard
|
|
K&A S4CL/S8CL
|
Luer
|
Barrel column with luer fitting at either end
|
Also known as Standard. For this instrument, we recommend the Luer (Standard) column as the ALL-FIT columns have a wider barrel.
|
|
Dr Oligo 48
|
MerMade
|
Pipette type column
|
A MerMade column is also known as a Supercolumn
|
|
Dr Oligo 192XLc, 768XLc just plates
|
MerMade, Syringe (up to 1.3 mL)
|
Pipette type column
|
A MerMade column is also known as a Supercolumn
|
|
OligoMaker X12, 48, 192, X192, X96
|
MerMade, Syringe (up to 1.3 mL)
|
Pipette type column
|
A MerMade column is also known as a Supercolumn
|