You are currently seeing list prices, to see your prices please log in
Baseline-ZERO DNase
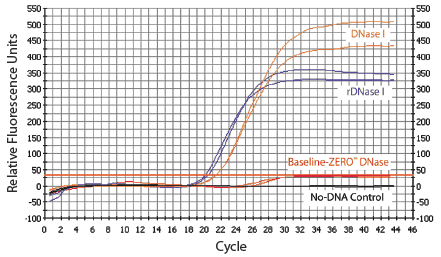
Digest large or small DNA molecules better than standard DNase I.
Key features
Show- Effectively digest large or small dsDNA and ssDNA into mononucleotides
- Digest unwanted dsDNA and ssDNA molecules including small ssDNA such as random hexamers
- Use in sensitive applications requiring reverse transcription where any contaminating DNA is unwanted
Size: 5000 units
Added to basket
Product information
SDS
Manuals and user guides
Product information sheets
Access support
Need some support with placing an order, setting up an account, or finding the right protocol?
Contact us