2-Aminopurine CE-Phosphoramidite
CAS No.:178925-41-2
Phosphoramidite for incorporation of a 2-amino-modified purine in a synthetic oligonucleotide.
Phosphoramidite for incorporation of a 2-amino-modified purine in a synthetic oligonucleotide.
Guanosine analogue phosphoramidite which lacks the O6 carbonyl_x000D_ resulting in a_x000D_ hydrogen bonding pattern isomeric_x000D_ with that of adenosine.
Phosphoramidite used to incorporate a 2-amino-modified deoxyadenosine into an oligonucleotide.
Explore our wide-ranging phosphoramidites and solid supports for generating oligos that help stabilize DNA duplexes, elucidate 3D structures of DNA, investigate DNA damage and epigenetic processes, and more. Several oligo modifications are useful for studying the structure, activity and function of DNA in various biological processes.
By partnering with LGC, Biosearch Technologies, you can get everything you need to synthesise modified oligos for your specific structural study.
We offer a wide portfolio of phosphoramidites and solid supports to help you:
We offer the Nap-dU CE-Phosphoramidite (NACN5-001) which can be used to incorporate functional diversity into oligonucleotides for aptamer research.
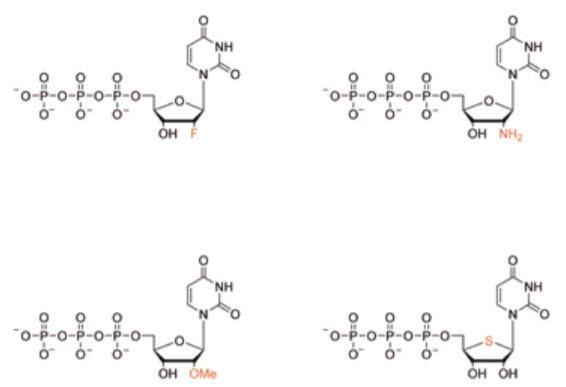
Figure 1: Chemical structures of sugar modified nucleotides used in the aptamer generation resulting in increased nuclease resistance : 2’-fluorouridine-5’-triphosphate, 2’-aminouridine-5’-triphosphate, 2’-methoxyuridine-5’-triphosphate and 4’-thiouridine-5’-triphosphate, These are currently not available from Biosearch Technologies as stock items. However, contact our Customer Service for a custom order (terms apply).
At Biosearch Technologies, we offer several amidites to incorporate hydroxy, hydroxymethyl, carboxy, and formyl modifications into your oligo for studying oxidative damage and repair, methylation and epigenetics.
Once incorporated into an oligonucleotide, these modifiers represent the various products in the biochemical pathway of dC modification.
Stabilizing duplexes is challenging with natural bases, so we offer several modifications—as phosphoramidites and CPG—that can be incorporated into your oligo at your desired location and for your specific application.
| Duplex Modifiers | Description |
|---|---|
| C-5 Methyl pyrimidine | Known to stabilise duplexes and enhance binding relative to the non- methylated bases.
|
| Amino phosphoramidites | One of the simplest methods of improving duplex stabilisation is the use of 2-Amino-dA CE Phosphoramidite (2,6-diaminopurine) in place of dA.
|
| Deoxyinosine (dI) | Often used as a degenerate base in an oligonucleotide to alleviate the problem of degeneracy of the genetic code.
|
| Deoxyuridine base | Used to induce mutagenic effects.
|
| Xanthosine (dX) | Considered a ‘universal base’ that can pair equally well with all four natural nucleosides.
|
We provide a wide range of halogenated nucleoside phosphoramidites and CPG supports, which can be used to synthesize oligos suitable for probing the three-dimensional structure of DNA by x-ray crystallography.
5-Fluoro-deoxyuridine is a base analogue that has the potential to bind to A and G without destabilising duplex formation, and it is an alternative to using mixed bases A/G for degeneracy.
2-Aminopurine-CE Phosphoramidite is useful for investigating structural changes, as the base is deficient in hydrogen bonding sites. It is also mildly fluorescent.
8-oxo-dG-CE Phosphoramidite can be sued to investigate the structure and activity of oligonucleotides containing an 8-oxo mutation. This is formed naturally when DNA is subjected to oxidative conditions or ionising radiation. The resulting 8-oxo modification is significant in mutagenesis and ultimately carcinogenesis.
| Europe, Middle East, and Africa | |
|---|---|
| UK | +44 1992 470 757 |
| Germany | +49 30 5304 2200 |
| North America, Latin America | |
| Wisconsin, USA | +1 888 575 9695 |
| Asia Pacific | |
| China | +8621-22509000 |
| Singapore | +65 6734 4800 |